LogosKG is the framework that combines matrix-based efficiency with billion-scale scalability and complete path reconstruction—all on standard hardware.
| Method (year) | Matrix-based | Scalability | Path Tracking | Device |
|---|---|---|---|---|
| Neo4j (Neo4j, Inc., 2025) | ✗ | ✓ | ✓ | CPU |
| TigerGraph (Deutsch et al., 2019) | ✗ | ✓ | ✓ | CPU |
| GraphBLAS (Kepner et al., 2016) | ✓ | ✗ | ✗ | CPU |
| igraph (Csardi and Nepusz, 2006) | ✗ | ✗ | ✓ | CPU |
| NetworkX (Hagberg et al., 2008) | ✗ | ✗ | ✓ | CPU |
| graph-tool (Peixoto, 2014) | ✗ | ✗ | ✓ | CPU |
| SNAP (Leskovec and Sosič, 2016) | ✗ | ✓ | ✓ | CPU |
| cuGraph (RAPIDS AI, 2025) | ✓ | ✗ | ✓ | GPU |
| DGL (Wang et al., 2019) | ✗ | ✓ | ✗ | GPU |
| PyG (Fey and Lenssen, 2019) | ✗ | ✓ | ✗ | GPU |
| LogosKG (ours, 2025) | ✓ | ✓ | ✓ | CPU/GPU |
Table 1: Comparison of retrieval systems and libraries. Matrix-based indicates the use of linear-algebra primitives for retrieval. Note: Entries reflect default design; unavailable features can be achieved with additional code or preprocessing, at the cost of extra time and memory.
The Challenge: Traditional graph retrieval methods struggle with deep multi-hop queries over large knowledge graphs. As query depth increases, memory usage explodes and response times become impractical, forcing researchers to limit exploration to just 1-3 hops, missing valuable distant connections.
Efficient
Matrix-based operations replace pointer-chasing, achieving sub-second multi-hop queries even on deep knowledge graph traversals.
Scalable
Integrates graph partitioning, cross-partition routing, and on-demand caching to enable querying of massive knowledge graphs on a single machine, even with limited hardware resources.
Interpretable
Preserves complete reasoning paths, allowing clinicians and researchers to trace exact multi-hop connections for transparent diagnosis.
Model Agnostic
Easily integrated as a plug-and-play retrieval module to enhance downstream LLMs and various clinical decision support applications.
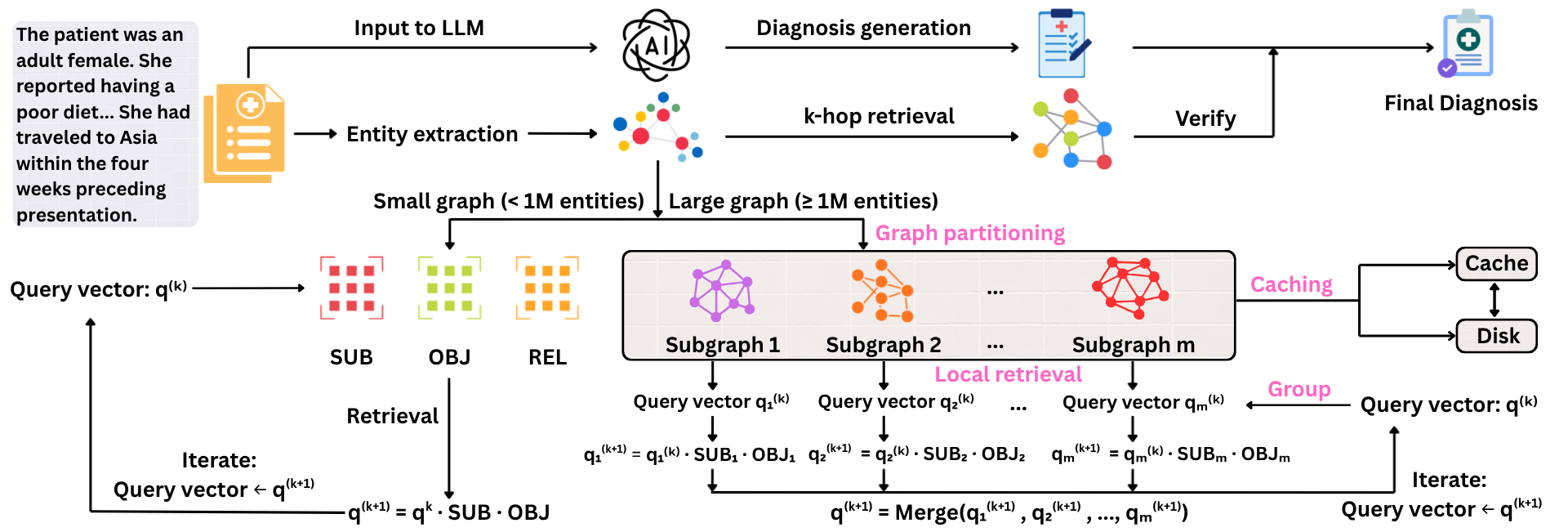
LogosKG uses a matrix-based workflow to process patient input, extract entities, and generate an initial diagnosis with an LLM. For very large graphs, it splits the graph into smaller subgraphs to achieve fast retrieval locally and merge the results globally. The retrieved evidence is then used to filter and enhance the diagnosis in a two-round pass, which improves prediction quality.
Comprehensive evaluation of retrieval fidelity, computational efficiency, scalability, and downstream clinical impact.
Retrieval Fidelity
We evaluate how accurately LogosKG retrieves relevant biomedical entities compared to established baselines using Jaccard similarity across hop distances.
Table A.1: Jaccard Similarity (LogosKG vs. Baselines)
| LogosKG family | Hop 1 | Hop 2 | Hop 3 | Hop 4 | Hop 5 |
|---|---|---|---|---|---|
| Neo4j | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| GraphBLAS | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| igraph | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| NetworkX | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| graph-tool | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| SNAP | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| cuGraph | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| DGL | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| PyG | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
Higher values indicate stronger agreement (1.00 = identical results). LogosKG maintains perfect fidelity with established CPU/GPU baselines.
Computational Efficiency
Comparison of Query Time (QT) in milliseconds and Timeout Rate (TR) across increasing hop distances on a standard CPU workload.
| Method | Hop 1 (2000 ms) | Hop 2 (4000 ms) | Hop 3 (6000 ms) | Hop 4 (8000 ms) | Hop 5 (10000 ms) | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| QT (ms) | TR (%) | QT (ms) | TR (%) | QT (ms) | TR (%) | QT (ms) | TR (%) | QT (ms) | TR (%) | |
| Baselines | ||||||||||
| NetworkX | 0.21 | 0.00 | 5.47 | 0.00 | 93.92 | 0.00 | 621.95 | 0.00 | 1511.28 | 0.00 |
| igraph | 1.15 | 0.00 | 26.13 | 0.00 | 309.90 | 0.00 | 837.12 | 0.00 | 580.91 | 0.00 |
| graph-tool | > 1458.79 | 45.33 | > 1900.01 | 10.00 | > 2141.41 | 2.00 | > 2396.95 | 0.67 | > 2306.34 | 0.67 |
| SNAP | 1.80 | 0.00 | 10.02 | 0.00 | 115.00 | 0.00 | 378.81 | 0.00 | 446.15 | 0.00 |
| GraphBLAS | 3.03 | 0.00 | 43.64 | 0.00 | 291.89 | 0.00 | 528.07 | 0.00 | 415.43 | 0.00 |
| Neo4j | > 923.86 | 11.33 | > 1946.43 | 20.00 | > 5739.02 | 95.33 | > 8000.00 | 100 | > 10000.00 | 100 |
| cuGraph | > 722.66 | 3.33 | > 919.30 | 0.67 | 1204.00 | 0.00 | 1504.82 | 0.00 | 1616.55 | 0.00 |
| DGL | > 966.65 | 10.00 | > 989.91 | 0.67 | 1042.25 | 0.00 | 1121.70 | 0.00 | 1099.26 | 0.00 |
| PyG | 249.66 | 0.00 | 271.46 | 0.00 | 365.90 | 0.00 | 646.96 | 0.00 | 735.34 | 0.00 |
| LogosKG family | ||||||||||
| LogosKG (Numba) | 12.28 | 0.00 | 28.72 | 0.00 | 77.65 | 0.00 | 140.07 | 0.00 | 204.25 | 0.00 |
| LogosKG (SciPy) | 13.81 | 0.00 | 34.97 | 0.00 | 104.21 | 0.00 | 289.61 | 0.00 | 677.23 | 0.00 |
| LogosKG (Torch-CPU) | 526.55 | 0.00 | 884.89 | 0.00 | 1321.05 | 0.00 | 1803.18 | 0.00 | 2207.25 | 0.00 |
| LogosKG (Torch-GPU) | 6.00 | 0.00 | 14.40 | 0.00 | 43.07 | 0.00 | 77.73 | 0.00 | 101.05 | 0.00 |
| LogosKG (Large-Numba) | 4.76 | 0.00 | 25.52 | 0.00 | 226.83 | 0.00 | 1411.72 | 0.00 | > 4085.72 | 4.00 |
| LogosKG (Large-SciPy) | 7.64 | 0.00 | 63.93 | 0.00 | 324.95 | 0.00 | > 1660.91 | 0.67 | > 4532.75 | 5.33 |
| LogosKG (Large-Torch-CPU) | 126.61 | 0.00 | > 1205.50 | 0.67 | > 2551.68 | 5.33 | > 4836.16 | 22.00 | > 7467.56 | 48.67 |
| LogosKG (Large-Torch-GPU) | 13.75 | 0.00 | 103.50 | 0.00 | 412.38 | 0.00 | 1634.93 | 0.00 | > 4482.14 | 6.00 |
Scalability Analysis
Performance analysis of LogosKG-Large on the PKG dataset across varying hops, batch sizes, cache sizes, and backends.
| Exp. 1: hops (Numba, cache size n = 16, batch size=50) | ||||
|---|---|---|---|---|
| Factor | Value | QT (ms) | Loads | Evicts |
| hops | 1 | 3410.90 | 16 | 0 |
| hops | 2 | 1610.78 | 16 | 0 |
| hops | 3 | 6114.69 | 16 | 0 |
| hops | 4 | 19592.08 | 16 | 0 |
| hops | 5 | 62726.25 | 16 | 0 |
| Exp. 2: batch size (Numba, cache size n = 16, hops k = 2) | ||||
|---|---|---|---|---|
| Factor | Value | QT (ms) | Loads | Evicts |
| batch size | 1 | 100912.09 | 12 | 0 |
| batch size | 10 | 5489.45 | 16 | 0 |
| batch size | 25 | 1365.09 | 16 | 0 |
| batch size | 50 | 1499.39 | 16 | 0 |
| batch size | 100 | 1444.68 | 16 | 0 |
| batch size | 150 | 1483.49 | 16 | 0 |
| Exp. 3: cache size (Numba, hops k = 2, batch size=50) | ||||
|---|---|---|---|---|
| Factor | Value | QT (ms) | Loads | Evicts |
| cache size | 1 | 441870.19 | 3010 | 3009 |
| cache size | 2 | 419436.59 | 2918 | 2916 |
| cache size | 4 | 384086.42 | 2659 | 2655 |
| cache size | 8 | 304066.00 | 1967 | 1959 |
| cache size | 16 | 4037.69 | 16 | 0 |
| Exp. 4: backend (cache size n = 16, hops k = 2, batch size=50) | ||||
|---|---|---|---|---|
| Factor | Value | QT (ms) | Loads | Evicts |
| backend | Numba | 4143.16 | 16 | 0 |
| backend | SciPy | 3889.86 | 16 | 0 |
| backend | Torch-CPU | 245311.21 | 16 | 0 |
| backend | Torch-GPU | 6409.32 | 16 | 0 |
Featured Projects
Clinical Diagnosis Prediction
Leveraging LogosKG to enhance LLM diagnostic accuracy on complex patient narratives through 2-round multi-hop retrieval.
KG-Based LLM Finetuning
Fine-tuning large language models with structured reasoning paths retrieved by LogosKG to improve downstream applications, such as biomedical reasoning.
Biomedical Discovery
Scaling retrieval across billion-scale biomedical knowledge graphs for robust hypothesis generation and validation.
